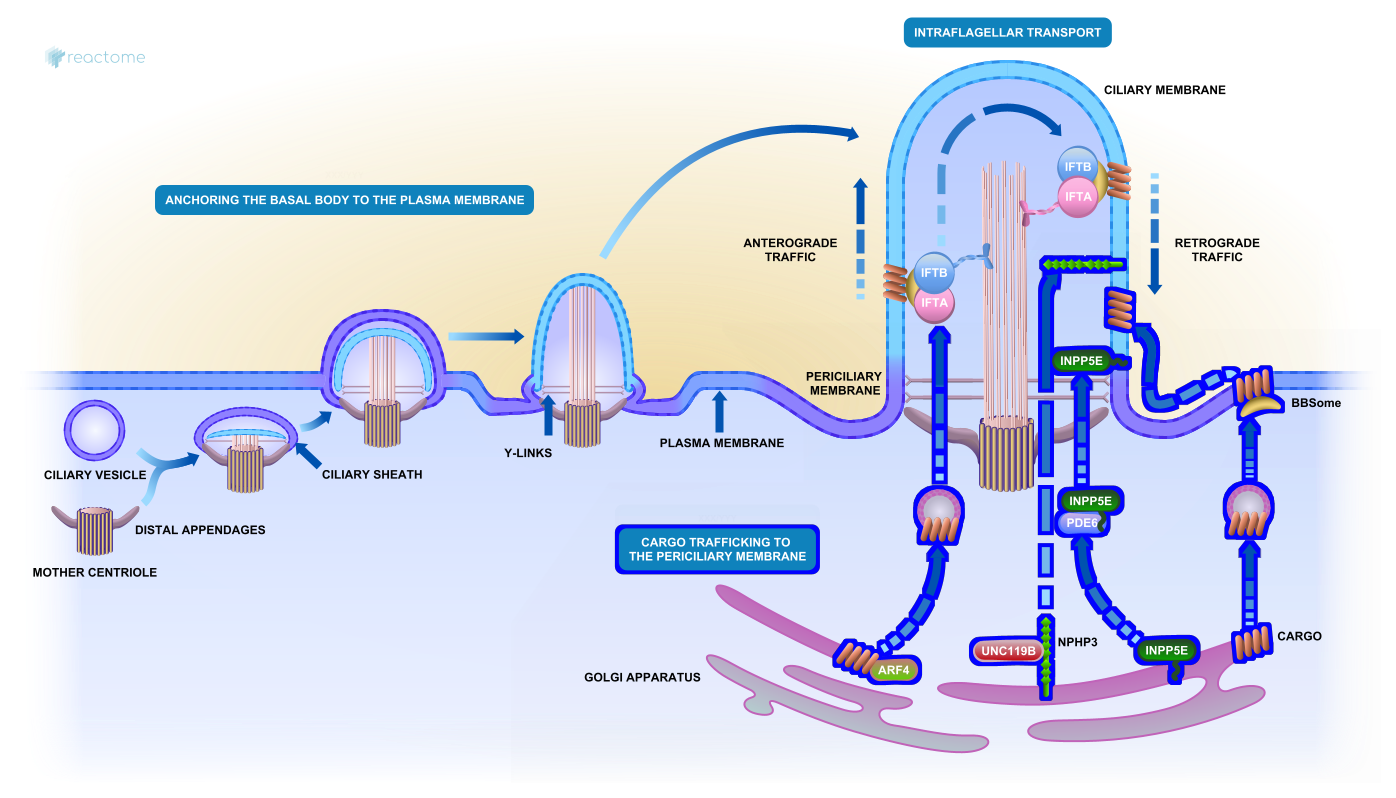
Assembly of the 9+0 primary cilium
New and Updated Topics and Pathways. Topics with new or revised pathways in this release include Cell-cell communication (Activation of STAT3 by cadherin engagement), Chromatin organization (CHD chromatin remodelers), Developmental biology (Differentiation of naive CD4+ T cells to T helper 2 cells (Th2 cells)), Disease (Defects of coagulation cascade) and Defects of contact activation system), DNA repair (HDR through Single Strand Annealing (SSA)), Gene expression (PRC2 methylates histones and DNA), Hemostasis (Coagulation pathway), Immune system (FXII activates plasma kallikrein-kinin system and FXIIa, PKa-dependent activation of coagulation pathway, Regulation of Complement cascade), Metabolism (choline metabolism, Inositol phosphate metabolism and Thyroxine metabolism and iodide transport), Muscle Contraction (Cardiac Conduction), Signal Transduction (Formation of the beta-catenin:TCF transactivating complex and MTOR signalling), and Transport of small molecules (Organic anion transport by SLC22 transporters).
New and Updated Illustrations. New or revised illustrations with embedded navigation features have been created for Cilium assembly, Assembly of the 9+0 primary cilium, ATP-dependent chromatin remodelers, Defects of coagulation cascade, Defects of contact activation system and kallikrein-kinin system, Differentiation of T cells, Diseases of hemostasis, Diseases of immune system, Hemostasis, and Innate immune system.
Thanks to our Contributors. Hanad Adan, Juliet Daniel, Anthony J Gesino, Leda Raptis, and Arielle Vaglio are our external authors and Isabella Barbutti, David P Hill, Lin Huang, Graham M Lord, are Alvin H Schmaier our external reviewers.
Annotation Statistics. Reactome comprises 16,338 human reactions organized into 2,870 pathways involving 32,399 proteins and modified forms of proteins encoded by 11,452 different human genes, 16,145 complexes, 2,198 small molecules, and 1,102 drugs. These annotations are supported by 42,784 literature references. We have projected these reactions onto 83,074 orthologous proteins, creating 20,616 orthologous pathways in 14 non-human species. Version 96 has annotations for 5752 protein variants (mutated proteins) and their post-translationally modified forms, derived from 400 proteins, which have contributed to the annotation of 2056 disease-specific reactions and 788 pathways. Our Illustration library now includes over 2,500 icons and over 200 high level interactive pathway Illustrations.
Other news:
Tools and Data. Our services and software tools are designed for biologists, bioinformaticians, and software developers. Pathway data is available to view in our Pathway Browser, to analyze your own dataset, to download, and access programmatically through our Content and Analysis Services. The Reactome FIViz app and ReactomeGSA package provide tools for multi-omics data analysis. The idg.reactome.org Web Portal provides a collection of web-based tools to help researchers place understudied proteins in a pathway context.
Reactome is excited to announce the beta release of our redesigned Pathway Browser. This major update delivers a modernized user interface (UI) and user experience (UX), along with new visualization and analysis features. Please provide us with your feedback.
Documentation and Training. Visit our online User Guide to access documentation supporting pathway analysis of experimental data. The Developer's Zone provides detailed documentation regarding our software, tools, and web services. Training and learning materials can be found here.
About the Reactome Project. Reactome is a collaboration between groups at the Ontario Institute for Cancer Research, Oregon Health and Science University, New York University Langone Medical Center, and The EMBL - European Bioinformatics Institute. Reactome is both an ELIXIR Core Data Resource and a Global Core Biodata Resource and has been certified as a Trustworthy Data Repository by the CoreTrustSeal Standards and Certification Board.
Reactome annotation files and interaction data derived from Reactome are distributed under a Creative Commons Public Domain (CC0 1.0 Universal) Licence. A Creative Commons Attribution 4.0 International (CC BY 4.0) License applies to all software and code, database data dumps, and Pathway Illustrations (Enhanced High-Level Diagrams), Icon Library, Art, and Branding Materials. A full description of the new and updated content is available on the Reactome website.
Follow us on This email address is being protected from spambots. You need JavaScript enabled to view it. get frequent updates about new and updated pathways, feature updates, and more!
For more information: If you have a question, want to provide feedback, or are interested in collaborating with us to annotate a topic, please contact us at This email address is being protected from spambots. You need JavaScript enabled to view it..

